7 DNA and RNA
Nucleic acids are biopolymers that are essential to all known forms of life. The term nucleic acid is the overall name for DNA and RNA. They are composed of nucleotides, which are the monomers made of three components: a 5-carbon sugar, a phosphate group and a nitrogenous base. If the sugar is a compound ribose, the polymer is RNA (ribonucleic acid); if the sugar is derived from ribose as deoxyribose, the polymer is DNA (deoxyribonucleic acid).
7.1 DNA
Deoxyribonucleic acid (DNA) is a molecule composed of two chains that coil around each other to form a double helix carrying genetic instructions for the development, functioning, growth and reproduction of all known organisms and many viruses. DNA and ribonucleic acid (RNA) are nucleic acids; alongside proteins, lipids and complex carbohydrates (polysaccharides), nucleic acids are one of the four major types of macromolecules that are essential for all known forms of life.
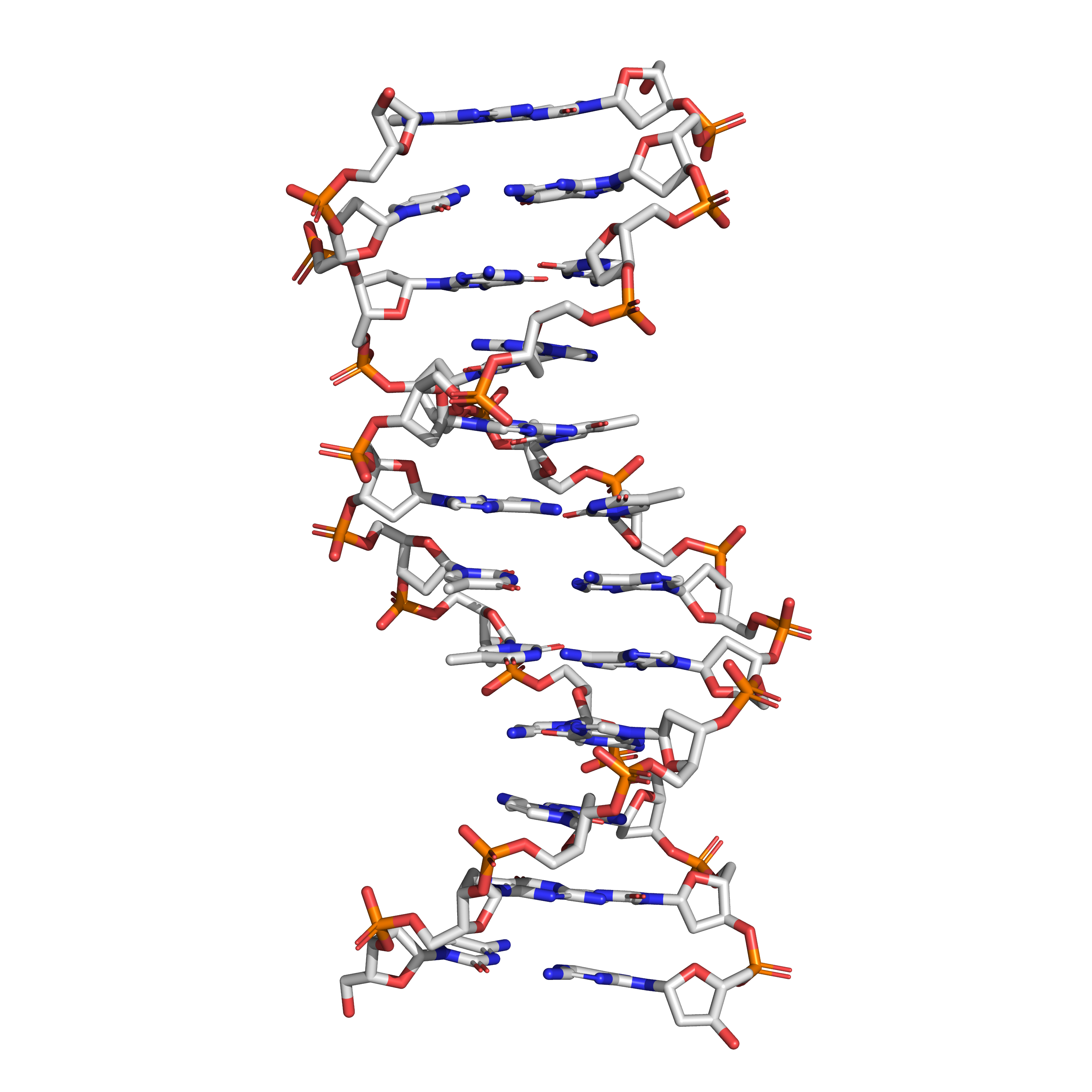
Figure 7.1: The structure of the DNA double helix. A section of DNA. The bases lie horizontally between the two spiraling strands. The atoms in the structure are colour-coded by element (based on atomic coordinates of PDB 1bna rendered with open source molecular visualization tool PyMol.)
The two DNA strands are also known as polynucleotides as they are composed of simpler monomeric units called nucleotides. Each nucleotide is composed of one of four nitrogen-containing nucleobases (cytosine [C], guanine [G], adenine [A] or thymine [T]), a sugar called deoxyribose, and a phosphate group. The nucleotides are joined to one another in a chain by covalent bonds between the sugar of one nucleotide and the phosphate of the next, resulting in an alternating sugar-phosphate backbone. The nitrogenous bases of the two separate polynucleotide strands are bound together, according to base pairing rules (A with T and C with G), with hydrogen bonds to make double-stranded DNA. The complementary nitrogenous bases are divided into two groups, pyrimidines and purines. In DNA, the pyrimidines are thymine and cytosine; the purines are adenine and guanine.
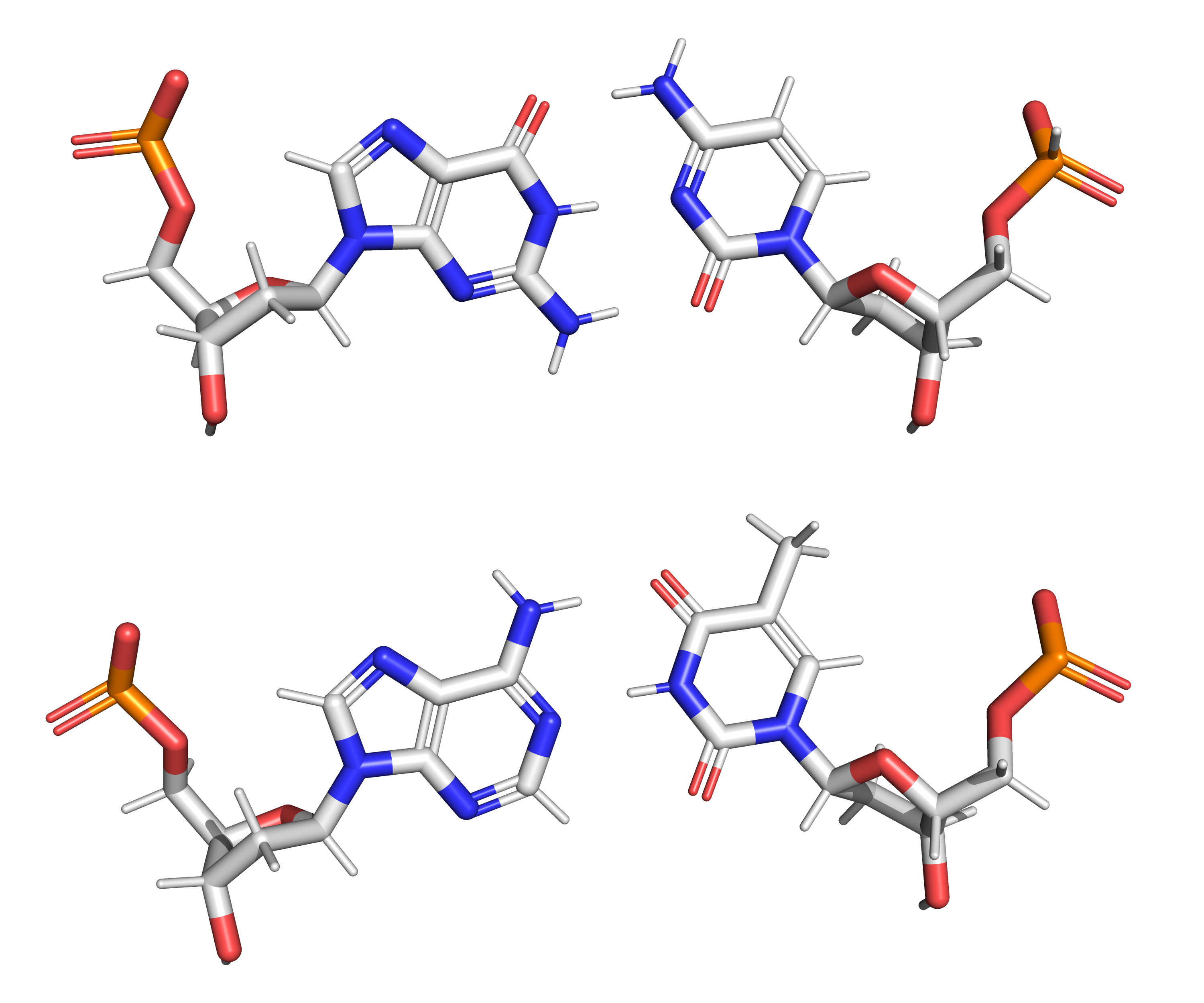
Figure 7.2: The structure of the four nucleotides and their base pairing in the DNA double helix. The atoms in the structure are colour-coded by element (based on atomic coordinates of PDB 1bna rendered with open source molecular visualization tool PyMol.)
Both strands of DNA store biological information. This information is replicated as and when the two strands separate. A large part of DNA (more than 98% for humans) is non-coding, meaning that these sections do not serve as patterns for protein sequences. The two strands of DNA run in opposite directions to each other and are thus antiparallel. Attached to each sugar is one of four types of nucleobases (informally, bases). It is the sequence of these four nucleobases along the backbone that encodes genetic information. RNA strands are created using DNA strands as a template in a process called transcription, where DNA bases are exchanged for their corresponding bases except in the case of thymine (T), which RNA substitutes for uracil (U). Under the genetic code, these RNA strands specify the sequence of amino acids within proteins in a process called translation.
Within eukaryotic cells, DNA is organized into long structures called chromosomes. Before typical cell division, these chromosomes are duplicated in the process of DNA replication, providing a complete set of chromosomes for each daughter cell. Eukaryotic organisms (animals, plants, fungi and protists) store most of their DNA inside the cell nucleus as nuclear DNA, and some in the mitochondria as mitochondrial DNA or in chloroplasts as chloroplast DNA. In contrast, prokaryotes (bacteria and archaea) store their DNA only in the cytoplasm, in circular chromosomes. Within eukaryotic chromosomes, chromatin proteins, such as histones, compact and organize DNA. These compacting structures guide the interactions between DNA and other proteins, helping control which parts of the DNA are transcribed.
DNA was first isolated by Friedrich Miescher in 1869. Its molecular structure was first identified by Francis Crick and James Watson at the Cavendish Laboratory within the University of Cambridge in 1953, whose model-building efforts were guided by X-ray diffraction data acquired by Raymond Gosling, who was a post-graduate student of Rosalind Franklin.
7.1.1 Structure and composition of DNA
DNA is a long polymer made from repeating units called nucleotides, each of which is usually symbolized by a single letter: either A, T, C, or G. The structure of DNA is dynamic along its length, being capable of coiling into tight loops and other shapes. In all species it is composed of two helical chains, bound to each other by hydrogen bonds. Both chains are coiled around the same axis, and have the same pitch of 34 angstroms (Å) (3.4 nanometres). The pair of chains has a radius of 10 angstroms (1.0 nanometre). Although each individual nucleotide is very small, a DNA polymer can be very large and contain hundreds of millions, such as in chromosome 1. Chromosome 1 is the largest human chromosome with approximately 220 million base pairs, and would be 85 mm long if straightened.
DNA does not usually exist as a single strand, but instead as a pair of strands that are held tightly together. These two long strands coil around each other, in the shape of a double helix. The nucleotide contains both a segment of the backbone of the molecule (which holds the chain together) and a nucleobase (which interacts with the other DNA strand in the helix). A nucleobase linked to a sugar is called a nucleoside, and a base linked to a sugar and to one or more phosphate groups is called a nucleotide. A biopolymer comprising multiple linked nucleotides (as in DNA) is called a polynucleotide.
The backbone of the DNA strand is made from alternating phosphate and sugar residues. The sugar in DNA is 2-deoxyribose, which is a pentose (five-carbon) sugar. The sugars are joined together by phosphate groups that form phosphodiester bonds between the third and fifth carbon atoms of adjacent sugar rings. These are known as the 3′-end (three prime end), and 5′-end (five prime end) carbons, the prime symbol being used to distinguish these carbon atoms from those of the base to which the deoxyribose forms a glycosidic bond. When imagining DNA, each phosphoryl is normally considered to “belong” to the nucleotide whose 5′ carbon forms a bond therewith. Any DNA strand therefore normally has one end at which there is a phosphoryl attached to the 5′ carbon of a ribose (the 5′ phosphoryl) and another end at which there is a free hydroxyl attached to the 3′ carbon of a ribose (the 3′ hydroxyl). The orientation of the 3′ and 5′ carbons along the sugar-phosphate backbone confers directionality (sometimes called polarity) to each DNA strand. In a nucleic acid double helix, the direction of the nucleotides in one strand is opposite to their direction in the other strand: the strands are antiparallel. The asymmetric ends of DNA strands are said to have a directionality of five prime end (5′ ), and three prime end (3′), with the 5′ end having a terminal phosphate group and the 3′ end a terminal hydroxyl group. One major difference between DNA and RNA is the sugar, with the 2-deoxyribose in DNA being replaced by the alternative pentose sugar ribose in RNA.
7.1.1.1 Grooves
Twin helical strands form the DNA backbone. Another double helix may be found tracing the spaces, or grooves, between the strands (Figure 7.3). These voids are adjacent to the base pairs and may provide a binding site. As the strands are not symmetrically located with respect to each other, the grooves are unequally sized. One groove, the major groove, is 22 angstroms (Å) wide and the other, the minor groove, is 12 Å wide. The width of the major groove means that the edges of the bases are more accessible in the major groove than in the minor groove. As a result, proteins such as transcription factors that can bind to specific sequences in double-stranded DNA usually make contact with the sides of the bases exposed in the major groove.

Figure 7.3: DNA major and minor grooves. PDB 1bna rendered with open source molecular visualization tool PyMol.)
7.1.1.2 Base pairing
In a DNA double helix, each type of nucleobase on one strand bonds with just one type of nucleobase on the other strand. This is called complementary base pairing. Here, purines form hydrogen bonds to pyrimidines, with adenine bonding only to thymine in two hydrogen bonds, and cytosine bonding only to guanine in three hydrogen bonds (Figure 7.4). This arrangement of two nucleotides binding together across the double helix is called a Watson-Crick base pair. As hydrogen bonds are not covalent, they can be broken and rejoined relatively easily. The two strands of DNA in a double helix can thus be pulled apart like a zipper, either by a mechanical force or high temperature. As a result of this base pair complementarity, all the information in the double-stranded sequence of a DNA helix is duplicated on each strand, which is vital in DNA replication. This reversible and specific interaction between complementary base pairs is critical for all the functions of DNA in organisms.
Figure 7.4: Top, a GC base pair with three hydrogen bonds. Bottom, an AT base pair with two hydrogen bonds. Non-covalent hydrogen bonds between the pairs are shown as dashed lines. The two types of base pairs form different numbers of hydrogen bonds, AT forming two hydrogen bonds, and GC forming three hydrogen bonds (see figures, right). DNA with high GC-content is more stable than DNA with low GC-content.
As noted above, most DNA molecules are actually two polymer strands, bound together in a helical fashion by noncovalent bonds; this double-stranded (dsDNA) structure is maintained largely by the intrastrand base stacking interactions, which are strongest for G,C stacks. The two strands can come apart—a process known as melting—to form two single-stranded DNA (ssDNA) molecules. Melting occurs at high temperature, low salt and high pH (low pH also melts DNA, but since DNA is unstable due to acid depurination, low pH is rarely used).
The stability of the dsDNA form depends not only on the GC-content (% G,C basepairs) but also on sequence (since stacking is sequence specific) and also length (longer molecules are more stable). The stability can be measured in various ways; a common way is the “melting temperature”, which is the temperature at which 50% of the ds molecules are converted to ss molecules; melting temperature is dependent on ionic strength and the concentration of DNA. As a result, it is both the percentage of GC base pairs and the overall length of a DNA double helix that determines the strength of the association between the two strands of DNA. Long DNA helices with a high GC-content have stronger-interacting strands, while short helices with high AT content have weaker-interacting strands. In biology, parts of the DNA double helix that need to separate easily, such as the TATAAT Pribnow box in some promoters, tend to have a high AT content, making the strands easier to pull apart.
In the laboratory, the strength of this interaction can be measured by finding the temperature necessary to break the hydrogen bonds, their melting temperature (also called Tm value). When all the base pairs in a DNA double helix melt, the strands separate and exist in solution as two entirely independent molecules. These single-stranded DNA molecules have no single common shape, but some conformations are more stable than others.
7.1.1.3 Sense and antisense
A DNA sequence is called a “sense” sequence if it is the same as that of a messenger RNA copy that is translated into protein. The sequence on the opposite strand is called the “antisense” sequence. Both sense and antisense sequences can exist on different parts of the same strand of DNA (i.e. both strands can contain both sense and antisense sequences). In both prokaryotes and eukaryotes, antisense RNA sequences are produced, but the functions of these RNAs are not entirely clear. One proposal is that antisense RNAs are involved in regulating gene expression through RNA-RNA base pairing.
A few DNA sequences in prokaryotes and eukaryotes, and more in plasmids and viruses, blur the distinction between sense and antisense strands by having overlapping genes. In these cases, some DNA sequences do double duty, encoding one protein when read along one strand, and a second protein when read in the opposite direction along the other strand. In bacteria, this overlap may be involved in the regulation of gene transcription, while in viruses, overlapping genes increase the amount of information that can be encoded within the small viral genome.
7.1.1.4 Supercoiling
DNA can be twisted like a rope in a process called DNA supercoiling. With DNA in its “relaxed” state, a strand usually circles the axis of the double helix once every 10.4 base pairs, but if the DNA is twisted the strands become more tightly or more loosely wound. If the DNA is twisted in the direction of the helix, this is positive supercoiling, and the bases are held more tightly together. If they are twisted in the opposite direction, this is negative supercoiling, and the bases come apart more easily. In nature, most DNA has slight negative supercoiling that is introduced by enzymes called topoisomerases. These enzymes are also needed to relieve the twisting stresses introduced into DNA strands during processes such as transcription and DNA replication.
7.1.2 Base modifications and DNA packaging
The expression of genes is influenced by how the DNA is packaged in chromosomes, in a structure called chromatin. Base modifications can be involved in packaging, with regions that have low or no gene expression usually containing high levels of methylation of cytosine bases. DNA packaging and its influence on gene expression can also occur by covalent modifications of the histone protein core around which DNA is wrapped in the chromatin structure or else by remodeling carried out by chromatin remodeling complexes. There is, further, crosstalk between DNA methylation and histone modification, so they can coordinately affect chromatin and gene expression.
For one example, cytosine methylation produces 5-methylcytosine, which is important for X-inactivation of chromosomes. The average level of methylation varies between organisms—the worm Caenorhabditis elegans lacks cytosine methylation, while vertebrates have higher levels, with up to 1% of their DNA containing 5-methylcytosine. Despite the importance of 5-methylcytosine, it can deaminate to leave a thymine base, so methylated cytosines are particularly prone to mutations. Other base modifications include adenine methylation in bacteria, the presence of 5-hydroxymethylcytosine in the brain, and the glycosylation of uracil to produce the “J-base” in kinetoplastids.
7.1.3 DNA damage
DNA can be damaged by many sorts of mutagens, which change the DNA sequence. Mutagens include oxidizing agents, alkylating agents and also high-energy electromagnetic radiation such as ultraviolet light and X-rays. The type of DNA damage produced depends on the type of mutagen. For example, UV light can damage DNA by producing thymine dimers, which are cross-links between pyrimidine bases. On the other hand, oxidants such as free radicals or hydrogen peroxide produce multiple forms of damage, including base modifications, particularly of guanosine, and double-strand breaks. A typical human cell contains about 150,000 bases that have suffered oxidative damage. Of these oxidative lesions, the most dangerous are double-strand breaks, as these are difficult to repair and can produce point mutations, insertions, deletions from the DNA sequence, and chromosomal translocations. These mutations can cause cancer. DNA damage that is naturally occurring, due to normal cellular processes that produce reactive oxygen species, the hydrolytic activities of cellular water, etc., also occurs frequently. Although most of this damage is repaired, in any cell some DNA damage may remain despite the action of repair processes. This DNA damage accumulates with age in mammalian postmitotic tissues. This accumulation appears to be an important underlying cause of aging.
Many mutagens fit into the space between two adjacent base pairs, this is called intercalation. Most intercalators are aromatic and planar molecules; examples include ethidium bromide, acridines, daunomycin, and doxorubicin. For an intercalator to fit between base pairs, the bases must separate, distorting the DNA strands by unwinding of the double helix. This inhibits both transcription and DNA replication, causing toxicity and mutations. As a result, DNA intercalators may be carcinogens, and in the case of thalidomide, a teratogen. Others such as benzo[a]pyrene diol epoxide and aflatoxin form DNA adducts that induce errors in replication. Nevertheless, due to their ability to inhibit DNA transcription and replication, other similar toxins are also used in chemotherapy to inhibit rapidly growing cancer cells.
7.1.4 Biological functions of DNA
DNA usually occurs as linear chromosomes in eukaryotes, and circular chromosomes in prokaryotes. The set of chromosomes in a cell makes up its genome; the human genome has approximately 3 billion base pairs of DNA arranged into 46 chromosomes. Transmission of genetic information in genes is achieved via complementary base pairing. For example, in transcription, the DNA sequence is copied into a complementary RNA sequence. Usually, this RNA copy is then used to make a matching protein sequence in a process called translation. In alternative fashion, a cell may simply copy its genetic information in a process called DNA replication.
7.1.4.1 Genes and genomes
Genomic DNA is tightly and orderly packed in the process called DNA condensation, to fit the small available volumes of the cell. In eukaryotes, DNA is located in the cell nucleus, with small amounts in mitochondria and chloroplasts. In prokaryotes, the DNA is held within an irregularly shaped body in the cytoplasm called the nucleoid.
In many species, only a small fraction of the total sequence of the genome encodes protein. For example, only about 1.5% of the human genome consists of protein-coding exons, with over 50% of human DNA consisting of non-coding repetitive sequences. The reasons for the presence of so much noncoding DNA in eukaryotic genomes and the extraordinary differences in genome size, or C-value, among species, represent a long-standing puzzle known as the “C-value enigma”. However, some DNA sequences that do not code protein may still encode functional non-coding RNA molecules, which are involved in the regulation of gene expression.
Some noncoding DNA sequences play structural roles in chromosomes. Telomeres and centromeres typically contain few genes but are important for the function and stability of chromosomes. An abundant form of noncoding DNA in humans are pseudogenes, which are copies of genes that have been disabled by mutation. These sequences are usually just molecular fossils, although they can occasionally serve as raw genetic material for the creation of new genes through the process of gene duplication and divergence.
7.1.4.2 Transcription and translation
A gene is a sequence of DNA that contains genetic information and can influence the phenotype of an organism. Within a gene, the sequence of bases along a DNA strand defines a messenger RNA sequence, which then defines one or more protein sequences. The relationship between the nucleotide sequences of genes and the amino-acid sequences of proteins is determined by the rules of translation, known collectively as the genetic code. The genetic code consists of three-letter ‘words’ called codons formed from a sequence of three nucleotides (e.g. ACT, CAG, TTT).
In transcription, the codons of a gene are copied into messenger RNA by RNA polymerase. This RNA copy is then decoded by a ribosome that reads the RNA sequence by base-pairing the messenger RNA to transfer RNA, which carries amino acids. Since there are 4 bases in 3-letter combinations, there are 64 possible codons. These encode the twenty standard amino acids, giving most amino acids more than one possible codon. There are also three ‘stop’ or ‘nonsense’ codons signifying the end of the coding region; these are the TAA, TGA, and TAG codons.
7.1.5 DNA replication
Cell division is essential for an organism to grow, but, when a cell divides, it must replicate the DNA in its genome so that the two daughter cells have the same genetic information as their parent. The double-stranded structure of DNA provides a simple mechanism for DNA replication. Here, the two strands are separated and then each strand’s complementary DNA sequence is recreated by an enzyme called DNA polymerase. This enzyme makes the complementary strand by finding the correct base through complementary base pairing and bonding it onto the original strand. As DNA polymerases can only extend a DNA strand in a 5′ to 3′ direction, different mechanisms are used to copy the antiparallel strands of the double helix. In this way, the base on the old strand dictates which base appears on the new strand, and the cell ends up with a perfect copy of its DNA.
7.1.6 Interactions of DNA with proteins
All the functions of DNA depend on interactions with proteins. These protein interactions can be non-specific, or the protein can bind specifically to a single DNA sequence. Enzymes can also bind to DNA and of these, the polymerases that copy the DNA base sequence in transcription and DNA replication are particularly important.
7.1.6.1 DNA-binding proteins
Structural proteins that bind DNA are well-understood examples of non-specific DNA-protein interactions. Within chromosomes, DNA is held in complexes with structural proteins. These proteins organize the DNA into a compact structure called chromatin. In eukaryotes, this structure involves DNA binding to a complex of small basic proteins called histones, while in prokaryotes multiple types of proteins are involved. The histones form a disk-shaped complex called a nucleosome, which contains two complete turns of double-stranded DNA wrapped around its surface. These non-specific interactions are formed through basic residues in the histones, making ionic bonds to the acidic sugar-phosphate backbone of the DNA, and are thus largely independent of the base sequence. Chemical modifications of these basic amino acid residues include methylation, phosphorylation, and acetylation. These chemical changes alter the strength of the interaction between the DNA and the histones, making the DNA more or less accessible to transcription factors and changing the rate of transcription. Other non-specific DNA-binding proteins in chromatin include the high-mobility group proteins, which bind to bent or distorted DNA. These proteins are important in bending arrays of nucleosomes and arranging them into the larger structures that make up chromosomes.
A distinct group of DNA-binding proteins is the DNA-binding proteins that specifically bind single-stranded DNA. In humans, replication protein A is the best-understood member of this family and is used in processes where the double helix is separated, including DNA replication, recombination, and DNA repair. These binding proteins seem to stabilize single-stranded DNA and protect it from forming stem-loops or being degraded by nucleases.
In contrast, other proteins have evolved to bind to particular DNA sequences. The most intensively studied of these are the various transcription factors, which are proteins that regulate transcription. Each transcription factor binds to one particular set of DNA sequences and activates or inhibits the transcription of genes that have these sequences close to their promoters. The transcription factors do this in two ways. Firstly, they can bind the RNA polymerase responsible for transcription, either directly or through other mediator proteins; this locates the polymerase at the promoter and allows it to begin transcription. Alternatively, transcription factors can bind enzymes that modify the histones at the promoter. This changes the accessibility of the DNA template to the polymerase.
As these DNA targets can occur throughout an organism’s genome, changes in the activity of one type of transcription factor can affect thousands of genes. Consequently, these proteins are often the targets of the signal transduction processes that control responses to environmental changes or cellular differentiation and development. The specificity of these transcription factors’ interactions with DNA come from the proteins making multiple contacts to the edges of the DNA bases, allowing them to “read” the DNA sequence. Most of these base-interactions are made in the major groove, where the bases are most accessible.
7.1.7 DNA-modifying enzymes
7.1.7.1 Nucleases and ligases
Nucleases are enzymes that cut DNA strands by catalyzing the hydrolysis of the phosphodiester bonds. Nucleases that hydrolyse nucleotides from the ends of DNA strands are called exonucleases, while endonucleases cut within strands. The most frequently used nucleases in molecular biology are the restriction endonucleases, which cut DNA at specific sequences. For instance, the EcoRI enzyme recognizes the 6-base sequence 5′-GAATTC-3′ and cuts each strand after the G creating 4 nucleotide sticky ends with a 5’ end overhang of AATT. In nature, these enzymes protect bacteria against phage infection by digesting the phage DNA when it enters the bacterial cell, acting as part of the restriction modification system. These sequence-specific nucleases are used in molecular cloning and DNA fingerprinting.
Enzymes called DNA ligases can rejoin cut or broken DNA strands. Ligases are particularly important in lagging strand DNA replication, as they join together the short segments of DNA produced at the replication fork into a complete copy of the DNA template. They are also used in DNA repair and genetic recombination.
7.1.7.2 Topoisomerases and helicases
Topoisomerases are enzymes with both nuclease and ligase activity. These proteins change the amount of supercoiling in DNA. Some of these enzymes work by cutting the DNA helix and allowing one section to rotate, thereby reducing its level of supercoiling; the enzyme then seals the DNA break. Other types of these enzymes are capable of cutting one DNA helix and then passing a second strand of DNA through this break, before rejoining the helix. Topoisomerases are required for many processes involving DNA, such as DNA replication and transcription.
Helicases are proteins that are a type of molecular motor. They use the chemical energy in nucleoside triphosphates, predominantly adenosine triphosphate (ATP), to break hydrogen bonds between bases and unwind the DNA double helix into single strands. These enzymes are essential for most processes where enzymes need to access the DNA bases.
7.1.7.3 Polymerases
Polymerases are enzymes that synthesize polynucleotide chains from nucleoside triphosphates. The sequence of their products is created based on existing polynucleotide chains—which are called templates. These enzymes function by repeatedly adding a nucleotide to the 3′ hydroxyl group at the end of the growing polynucleotide chain. As a consequence, all polymerases work in a 5′ to 3′ direction. In the active site of these enzymes, the incoming nucleoside triphosphate base-pairs to the template: this allows polymerases to accurately synthesize the complementary strand of their template. Polymerases are classified according to the type of template that they use.
In DNA replication, DNA-dependent DNA polymerases make copies of DNA polynucleotide chains. To preserve biological information, it is essential that the sequence of bases in each copy are precisely complementary to the sequence of bases in the template strand. Many DNA polymerases have a proofreading activity. Here, the polymerase recognizes the occasional mistakes in the synthesis reaction by the lack of base pairing between the mismatched nucleotides. If a mismatch is detected, a 3′ to 5′ exonuclease activity is activated and the incorrect base removed. In most organisms, DNA polymerases function in a large complex called the replisome that contains multiple accessory subunits, such as the DNA clamp or helicases.
RNA-dependent DNA polymerases are a specialized class of polymerases that copy the sequence of an RNA strand into DNA. They include reverse transcriptase, which is a viral enzyme involved in the infection of cells by retroviruses, and telomerase, which is required for the replication of telomeres. For example, HIV reverse transcriptase is an enzyme for AIDS virus replication. Telomerase is an unusual polymerase because it contains its own RNA template as part of its structure. It synthesizes telomeres at the ends of chromosomes. Telomeres prevent fusion of the ends of neighboring chromosomes and protect chromosome ends from damage.
Transcription is carried out by a DNA-dependent RNA polymerase that copies the sequence of a DNA strand into RNA. To begin transcribing a gene, the RNA polymerase binds to a sequence of DNA called a promoter and separates the DNA strands. It then copies the gene sequence into a messenger RNA transcript until it reaches a region of DNA called the terminator, where it halts and detaches from the DNA. As with human DNA-dependent DNA polymerases, RNA polymerase II, the enzyme that transcribes most of the genes in the human genome, operates as part of a large protein complex with multiple regulatory and accessory subunits.
7.1.8 DNA recombination
A DNA helix usually does not interact with other segments of DNA, and in human cells, the different chromosomes even occupy separate areas in the nucleus called “chromosome territories”. This physical separation of different chromosomes is important for the ability of DNA to function as a stable repository for information, as one of the few times chromosomes interact is in chromosomal crossover which occurs during sexual reproduction, when genetic recombination occurs. Chromosomal crossover is when two DNA helices break, swap a section and then rejoin.
Recombination allows chromosomes to exchange genetic information and produces new combinations of genes, which increases the efficiency of natural selection and can be important in the rapid evolution of new proteins. Genetic recombination can also be involved in DNA repair, particularly in the cell’s response to double-strand breaks.
The most common form of chromosomal crossover is homologous recombination, where the two chromosomes involved share very similar sequences. Non-homologous recombination can be damaging to cells, as it can produce chromosomal translocations and genetic abnormalities. The recombination reaction is catalyzed by enzymes known as recombinases, such as RAD51. The first step in recombination is a double-stranded break caused by either an endonuclease or damage to the DNA. A series of steps catalyzed in part by the recombinase then leads to joining of the two helices by at least one Holliday junction, in which a segment of a single strand in each helix is annealed to the complementary strand in the other helix. The Holliday junction is a tetrahedral junction structure that can be moved along the pair of chromosomes, swapping one strand for another. The recombination reaction is then halted by cleavage of the junction and re-ligation of the released DNA. Only strands of like polarity exchange DNA during recombination. There are two types of cleavage: east-west cleavage and north-south cleavage. The north-south cleavage nicks both strands of DNA, while the east-west cleavage has one strand of DNA intact.
7.2 Evolution
DNA contains the genetic information that allows all forms of life to function, grow and reproduce. However, it is unclear how long in the 4-billion-year history of life DNA has performed this function, as it has been proposed that the earliest forms of life may have used RNA as their genetic material. RNA may have acted as the central part of early cell metabolism as it can both transmit genetic information and carry out catalysis as part of ribozymes. This ancient RNA world where nucleic acid would have been used for both catalysis and genetics may have influenced the evolution of the current genetic code based on four nucleotide bases. This would occur, since the number of different bases in such an organism is a trade-off between a small number of bases increasing replication accuracy and a large number of bases increasing the catalytic efficiency of ribozymes. However, there is no direct evidence of ancient genetic systems, as recovery of DNA from most fossils is impossible because DNA survives in the environment for less than one million years, and slowly degrades into short fragments in solution.
Building blocks of DNA (adenine, guanine, and related organic molecules) may have been formed extraterrestrially in outer space. Complex DNA and RNA organic compounds of life, including uracil, cytosine, and thymine, have also been formed in the laboratory under conditions mimicking those found in outer space, using starting chemicals, such as pyrimidine, found in meteorites. Pyrimidine, like polycyclic aromatic hydrocarbons (PAHs), the most carbon-rich chemical found in the universe, may have been formed in red giants or in interstellar cosmic dust and gas clouds.
7.3 Genetic engineering
Methods have been developed to purify DNA from organisms, such as phenol-chloroform extraction, and to manipulate it in the laboratory, such as restriction digests and the polymerase chain reaction. Modern biology and biochemistry make intensive use of these techniques in recombinant DNA technology. Recombinant DNA is a man-made DNA sequence that has been assembled from other DNA sequences. They can be transformed into organisms in the form of plasmids or in the appropriate format, by using a viral vector. The genetically modified organisms produced can be used to produce products such as recombinant proteins, used in medical research, or be grown in agriculture.
7.4 DNA profiling
Forensic scientists can use DNA in blood, semen, skin, saliva or hair found at a crime scene to identify a matching DNA of an individual, such as a perpetrator. This process is formally termed DNA profiling, also called DNA fingerprinting. In DNA profiling, the lengths of variable sections of repetitive DNA, such as short tandem repeats and minisatellites, are compared between people. This method is usually an extremely reliable technique for identifying a matching DNA. However, identification can be complicated if the scene is contaminated with DNA from several people. DNA profiling was developed in 1984 by British geneticist Sir Alec Jeffreys, and first used in forensic science to convict Colin Pitchfork in the 1988 Enderby murders case.
DNA profiling is also used in DNA paternity testing to determine if someone is the biological parent or grandparent of a child with the probability of parentage is typically 99.99% when the alleged parent is biologically related to the child. Normally, paternity testing is performed after birth, but recently developed methods allow isolation and sequencing of fetal DNA from the blood of the mother.
7.5 RNA
Ribonucleic acid (RNA) is a polymeric molecule essential in various biological roles in coding, decoding, regulation and expression of genes. Like DNA, RNA is assembled as a chain of nucleotides, but unlike DNA it is more often found in nature as a single-strand folded onto itself, rather than a paired double-strand. Cellular organisms use messenger RNA (mRNA) to convey genetic information (using the nitrogenous bases of guanine, uracil, adenine, and cytosine, denoted by the letters G, U, A, and C) and direct synthesis of proteins by ribosomes. Many viruses encode their genetic information using an RNA genome.
Some RNA molecules play an active role within cells by catalyzing biological reactions, controlling gene expression, or sensing and communicating responses to cellular signals. One of these active processes is protein synthesis, a universal function in which RNA molecules direct the synthesis of proteins on ribosomes. This process uses transfer RNA (tRNA) molecules to deliver amino acids to the ribosome, where ribosomal RNA (rRNA) then links amino acids together to form coded proteins.
7.5.1 Comparison with DNA
The chemical structure of RNA is very similar to that of DNA, but differs in three primary ways:
- Unlike double-stranded DNA, RNA is a single-stranded molecule in many of its biological roles and consists of much shorter chains of nucleotides. However, a single RNA molecule can, by complementary base pairing, form intrastrand double helixes, as in tRNA.
- While the sugar-phosphate “backbone” of DNA contains deoxyribose, RNA contains ribose instead. Ribose has a hydroxyl group attached to the pentose ring in the 2’ position, whereas deoxyribose does not. The hydroxyl groups in the ribose backbone make RNA more chemically labile than DNA by lowering the activation energy of hydrolysis.
- The complementary base to adenine in DNA is thymine, whereas in RNA, it is uracil, which is an unmethylated form of thymine.
Like DNA, most biologically active RNAs, including mRNA, tRNA, rRNA, snRNAs, and other non-coding RNAs, contain self-complementary sequences that allow parts of the RNA to fold and pair with itself to form double helices. Analysis of these RNAs has revealed that they are highly structured. Unlike DNA, their structures do not consist of long double helices, but rather collections of short helices packed together into structures akin to proteins. In this fashion, RNAs can achieve chemical catalysis (like enzymes). For instance, determination of the structure of the ribosome—an RNA-protein complex that catalyzes peptide bond formation—revealed that its active site is composed entirely of RNA.
7.5.2 Structure of RNA
Each nucleotide in RNA contains a ribose sugar, with carbons numbered 1’ through 5’. A base is attached to the 1’ position, in general, adenine (A), cytosine (C), guanine (G), or uracil (U). Adenine and guanine are purines, cytosine and uracil are pyrimidines. A phosphate group is attached to the 3’ position of one ribose and the 5’ position of the next. The phosphate groups have a negative charge each, making RNA a charged molecule (polyanion). The bases form hydrogen bonds between cytosine and guanine, between adenine and uracil and between guanine and uracil. However, other interactions are possible, such as a group of adenine bases binding to each other in a bulge, or the GNRA tetraloop that has a guanine–adenine base-pair.
An important structural component of RNA that distinguishes it from DNA is the presence of a hydroxyl group at the 2’ position of the ribose sugar.
RNA is transcribed with only four bases (adenine, cytosine, guanine and uracil), but these bases and attached sugars can be modified in numerous ways as the RNAs mature. Pseudouridine (Ψ), in which the linkage between uracil and ribose is changed from a C–N bond to a C–C bond, and ribothymidine (T) are found in various places (the most notable ones being in the TΨC loop of tRNA). Another notable modified base is hypoxanthine, a deaminated adenine base whose nucleoside is called inosine (I). Inosine plays a key role in the wobble hypothesis of the genetic code.
There are more than 100 other naturally occurring modified nucleosides. The greatest structural diversity of modifications can be found in tRNA, while pseudouridine and nucleosides with 2’-O-methylribose often present in rRNA are the most common. The specific roles of many of these modifications in RNA are not fully understood. However, it is notable that, in ribosomal RNA, many of the post-transcriptional modifications occur in highly functional regions, such as the peptidyl transferase center and the subunit interface, implying that they are important for normal function.
The functional form of single-stranded RNA molecules, just like proteins, frequently requires a specific tertiary structure. The scaffold for this structure is provided by secondary structural elements that are hydrogen bonds within the molecule. This leads to several recognizable “domains” of secondary structure like hairpin loops, bulges, and internal loops. Since RNA is charged, metal ions such as Mg2+ are needed to stabilise many secondary and tertiary structures.
The naturally occurring enantiomer of RNA is D-RNA composed of D-ribonucleotides. All chirality centers are located in the D-ribose. By the use of L-ribose or rather L-ribonucleotides, L-RNA can be synthesized. L-RNA is much more stable against degradation by RNase.
7.5.3 Synthesis of RNA
Synthesis of RNA is usually catalyzed by an enzyme—RNA polymerase—using DNA as a template, a process known as transcription. Initiation of transcription begins with the binding of the enzyme to a promoter sequence in the DNA (usually found “upstream” of a gene). The DNA double helix is unwound by the helicase activity of the enzyme. The enzyme then progresses along the template strand in the 3’ to 5’ direction, synthesizing a complementary RNA molecule with elongation occurring in the 5’ to 3’ direction. The DNA sequence also dictates where termination of RNA synthesis will occur.
Primary transcript RNAs are often modified by enzymes after transcription. For example, a poly(A) tail and a 5’ cap are added to eukaryotic pre-mRNA and introns are removed by the spliceosome.
There are also a number of RNA-dependent RNA polymerases that use RNA as their template for synthesis of a new strand of RNA. For instance, a number of RNA viruses (such as poliovirus) use this type of enzyme to replicate their genetic material. Also, RNA-dependent RNA polymerase is part of the RNA interference pathway in many organisms.
7.5.4 Coding and non-coding RNA
Messenger RNA (mRNA) is the RNA that carries information from DNA to the ribosome, the sites of protein synthesis (translation) in the cell. The coding sequence of the mRNA determines the amino acid sequence in the protein that is produced. However, many RNAs do not code for protein (about 97% of the transcriptional output is non-protein-coding in eukaryotes).
These so-called non-coding RNAs (“ncRNA”) can be encoded by their own genes (RNA genes), but can also derive from mRNA introns. The most prominent examples of non-coding RNAs are transfer RNA (tRNA) and ribosomal RNA (rRNA), both of which are involved in the process of translation. There are also non-coding RNAs involved in gene regulation, RNA processing and other roles. Certain RNAs are able to catalyse chemical reactions such as cutting and ligating other RNA molecules, and the catalysis of peptide bond formation in the ribosome; these are known as ribozymes.
According to the length of RNA chain, RNA includes small RNA and long RNA. Usually, small RNAs are shorter than 200 nt in length, and long RNAs are greater than 200 nt long. Long RNAs, also called large RNAs, mainly include long non-coding RNA (lncRNA) and mRNA. Small RNAs mainly include 5.8S ribosomal RNA (rRNA), 5S rRNA, transfer RNA (tRNA), microRNA (miRNA), small interfering RNA (siRNA), small nucleolar RNA (snoRNAs), Piwi-interacting RNA (piRNA), tRNA-derived small RNA (tsRNA) and small rDNA-derived RNA (srRNA).
Messenger RNA (mRNA) carries genetic information from the nucleus to the cytoplasm and serves as template for the synthesis of protein by ribosomes, a process called translation. In eukaryotic cells, once precursor mRNA (pre-mRNA) has been transcribed from DNA, it is processed to mature mRNA. This removes its introns (non-coding stretches of DNA sequence) from the pre-mRNA. The mRNA is then exported from the nucleus to the cytoplasm, where ribosomes bind to it and translate its the corresponding protein form with the help of tRNA. In prokaryotic cells, which do not have nucleus and cytoplasm compartments, mRNA can bind to ribosomes while it is being transcribed from DNA. After a certain amount of time, the message degrades into its component nucleotides with the assistance of ribonucleases.
Transfer RNA (tRNA) is a small RNA chain of about 80 nucleotides that transfers a specific amino acid to a growing polypeptide chain at the ribosomal site of protein synthesis during translation. It has sites for amino acid attachment and an anticodon region for codon recognition that binds to a specific sequence on the messenger RNA chain through hydrogen bonding.

Figure 7.5: Tertiary structure of tRNA (based on atomic coordinates of PDB 1ehz rendered with open source molecular visualization tool PyMol.)
Ribosomal RNA (rRNA) is the catalytic component of the ribosomes. Eukaryotic ribosomes contain four different rRNA molecules: 18S, 5.8S, 28S and 5S rRNA. Three of the rRNA molecules are synthesized in the nucleolus, and one is synthesized elsewhere. In the cytoplasm, ribosomal RNA and protein combine to form a nucleoprotein called a ribosome. The ribosome binds mRNA and carries out protein synthesis. Several ribosomes may be attached to a single mRNA at any time. Nearly all the RNA found in a typical eukaryotic cell is rRNA.
Transfer-messenger RNA (tmRNA) is found in many bacteria and plastids. It tags proteins encoded by mRNAs that lack stop codons for degradation and prevents the ribosome from stalling.
7.5.5 Regulatory RNA
The earliest known regulators of gene expression were proteins known as repressors and activators, regulators with specific short binding sites within enhancer regions near the genes to be regulated. More recently, RNAs have been found to regulate genes as well. There are several kinds of RNA-dependent processes in eukaryotes regulating the expression of genes at various points, such as RNAi repressing genes post-transcriptionally, long non-coding RNAs shutting down blocks of chromatin epigenetically, and enhancer RNAs inducing increased gene expression. In addition to these mechanisms in eukaryotes, both bacteria and archaea have been found to use regulatory RNAs extensively. Bacterial small RNA and the CRISPR system are examples of such prokaryotic regulatory RNA systems.
7.5.6 RNA interference by miRNAs
Post-transcriptional expression levels of many genes can be controlled by RNA interference, in which miRNAs, specific short RNA molecules, pair with mRNA regions and target them for degradation. This antisense-based process involves steps that first process the RNA so that it can base-pair with a region of its target mRNAs. Once the base pairing occurs, other proteins direct the mRNA to be destroyed by nucleases.
Next to be linked to regulation were Xist and other long noncoding RNAs associated with X chromosome inactivation. Their roles, at first mysterious, were shown to be the silencing of blocks of chromatin via recruitment of Polycomb complex so that messenger RNA could not be transcribed from them. Additional lncRNAs, currently defined as RNAs of more than 200 base pairs that do not appear to have coding potential, have been found associated with regulation of stem cell pluripotency and cell division.
The third major group of regulatory RNAs is called enhancer RNAs. It is not clear at present whether they are a unique category of RNAs of various lengths or constitute a distinct subset of lncRNAs. In any case, they are transcribed from enhancers, which are known regulatory sites in the DNA near genes they regulate. They up-regulate the transcription of the gene(s) under control of the enhancer from which they are transcribed.
At first, regulatory RNA was thought to be a eukaryotic phenomenon, a part of the explanation for why so much more transcription in higher organisms was seen than had been predicted. But as soon as researchers began to look for possible RNA regulators in bacteria, they turned up there as well, termed as small RNA (sRNA). Bacterial small RNAs generally act via antisense pairing with mRNA to down-regulate its translation, either by affecting stability or affecting cis-binding ability. Riboswitches have also been discovered. They are cis-acting regulatory RNA sequences acting allosterically. They change shape when they bind metabolites so that they gain or lose the ability to bind chromatin to regulate expression of genes.
Archaea also have systems of regulatory RNA. The CRISPR system, recently being used to edit DNA in situ, acts via regulatory RNAs in archaea and bacteria to provide protection against virus invaders.
Many RNAs are involved in modifying other RNAs. Introns are spliced out of pre-mRNA by spliceosomes, which contain several small nuclear RNAs (snRNA), or the introns can be ribozymes that are spliced by themselves. RNA can also be altered by having its nucleotides modified to nucleotides other than A, C, G and U. In eukaryotes, modifications of RNA nucleotides are in general directed by small nucleolar RNAs (snoRNA; 60–300 nt), found in the nucleolus and cajal bodies. snoRNAs associate with enzymes and guide them to a spot on an RNA by basepairing to that RNA. These enzymes then perform the nucleotide modification. rRNAs and tRNAs are extensively modified, but snRNAs and mRNAs can also be the target of base modification. RNA can also be methylated.
RNA viruses have genomes composed of RNA that encodes a number of proteins. The viral genome is replicated by some of those proteins, while other proteins protect the genome as the virus particle moves to a new host cell. Viroids are another group of pathogens, but they consist only of RNA, do not encode any protein and are replicated by a host plant cell’s polymerase.
Reverse transcribing viruses replicate their genomes by reverse transcribing DNA copies from their RNA; these DNA copies are then transcribed to new RNA. Retrotransposons also spread by copying DNA and RNA from one another, and telomerase contains an RNA that is used as template for building the ends of eukaryotic chromosomes.
Research on RNA has led to many important biological discoveries and numerous Nobel Prizes. Nucleic acids were discovered in 1868 by Friedrich Miescher, who called the material ‘nuclein’ since it was found in the nucleus. It was later discovered that prokaryotic cells, which do not have a nucleus, also contain nucleic acids. The role of RNA in protein synthesis was suspected already in 1939. Severo Ochoa won the 1959 Nobel Prize in Medicine (shared with Arthur Kornberg) after he discovered an enzyme that can synthesize RNA in the laboratory. However, the enzyme discovered by Ochoa (polynucleotide phosphorylase) was later shown to be responsible for RNA degradation, not RNA synthesis. In 1956 Alex Rich and David Davies hybridized two separate strands of RNA to form the first crystal of RNA whose structure could be determined by X-ray crystallography.
The sequence of the 77 nucleotides of a yeast tRNA was found by Robert W. Holley in 1965, winning Holley the 1968 Nobel Prize in Medicine (shared with Har Gobind Khorana and Marshall Nirenberg).
During the early 1970s, retroviruses and reverse transcriptase were discovered, showing for the first time that enzymes could copy RNA into DNA (the opposite of the usual route for transmission of genetic information). For this work, David Baltimore, Renato Dulbecco and Howard Temin were awarded a Nobel Prize in 1975. In 1976, Walter Fiers and his team determined the first complete nucleotide sequence of an RNA virus genome, that of bacteriophage MS2.
In 1977, introns and RNA splicing were discovered in both mammalian viruses and in cellular genes, resulting in a 1993 Nobel to Philip Sharp and Richard Roberts. Catalytic RNA molecules (ribozymes) were discovered in the early 1980s, leading to a 1989 Nobel award to Thomas Cech and Sidney Altman. In 1990, it was found in Petunia that introduced genes can silence similar genes of the plant’s own, now known to be a result of RNA interference.
At about the same time, 22 nt long RNAs, now called microRNAs, were found to have a role in the development of C. elegans. Studies on RNA interference gleaned a Nobel Prize for Andrew Fire and Craig Mello in 2006, and another Nobel was awarded for studies on the transcription of RNA to Roger Kornberg in the same year. The discovery of gene regulatory RNAs has led to attempts to develop drugs made of RNA, such as siRNA, to silence genes. Adding to the Nobel prizes awarded for research on RNA in 2009 it was awarded for the elucidation of the atomic structure of the ribosome to Venki Ramakrishnan, Tom Steitz, and Ada Yonath.
In 1967, Carl Woese hypothesized that RNA might be catalytic and suggested that the earliest forms of life (self-replicating molecules) could have relied on RNA both to carry genetic information and to catalyze biochemical reactions—an RNA world.